Current research focus:
|
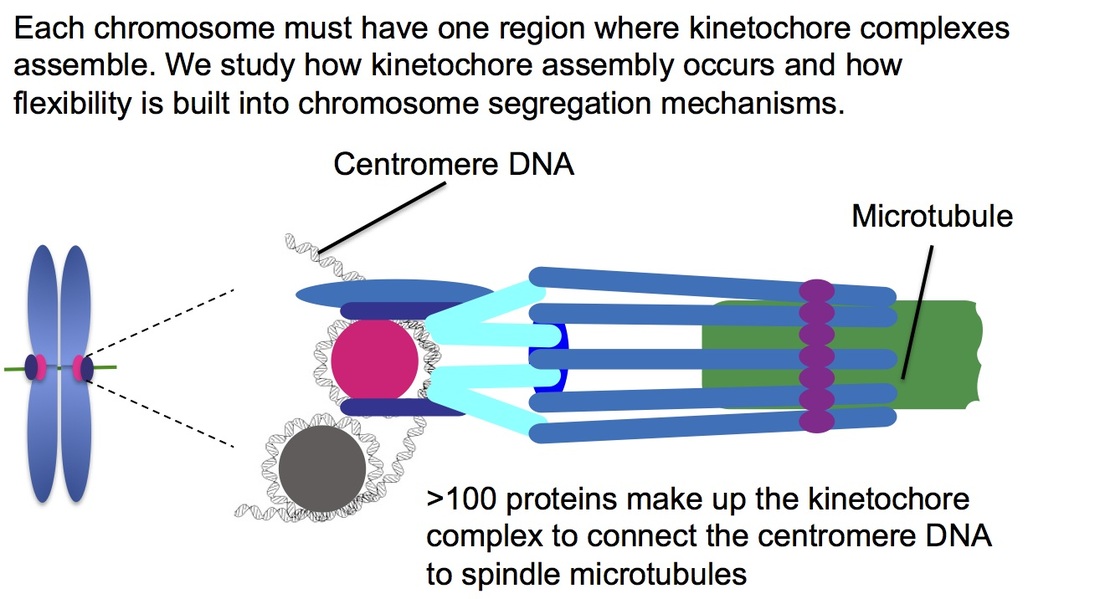
One aspect of biology that fascinates me is how organisms balance genome stability with adaptation to stress. If the balance tips toward genome stability, genetic diversity necessary for evolution will not arise. However, excess genome instability can be lethal and defects in genome stability are associated with a number of human diseases, including cancer. I am interested in how chromosome segregation mechanisms contribute to genome stability and how this process can be altered to allow the cell to adapt to changing conditions. The kinetochore is a large complex of proteins assembled at centromere DNA to allow attachment of the DNA to spindle microtubules and chromosome segregation during mitosis. In most eukaryotes, including humans, centromere regions are maintained epigenetically rather than being defined by a specific sequence and centromeres exhibit flexibility in size and position. In my research, I use the yeast Candida albicans as a model system to explore the mechanisms and consequences of centromere and kinetochore flexibility using a combination of molecular biology, microbiology, genetics, biochemistry, and cell biology techniques.
|
|
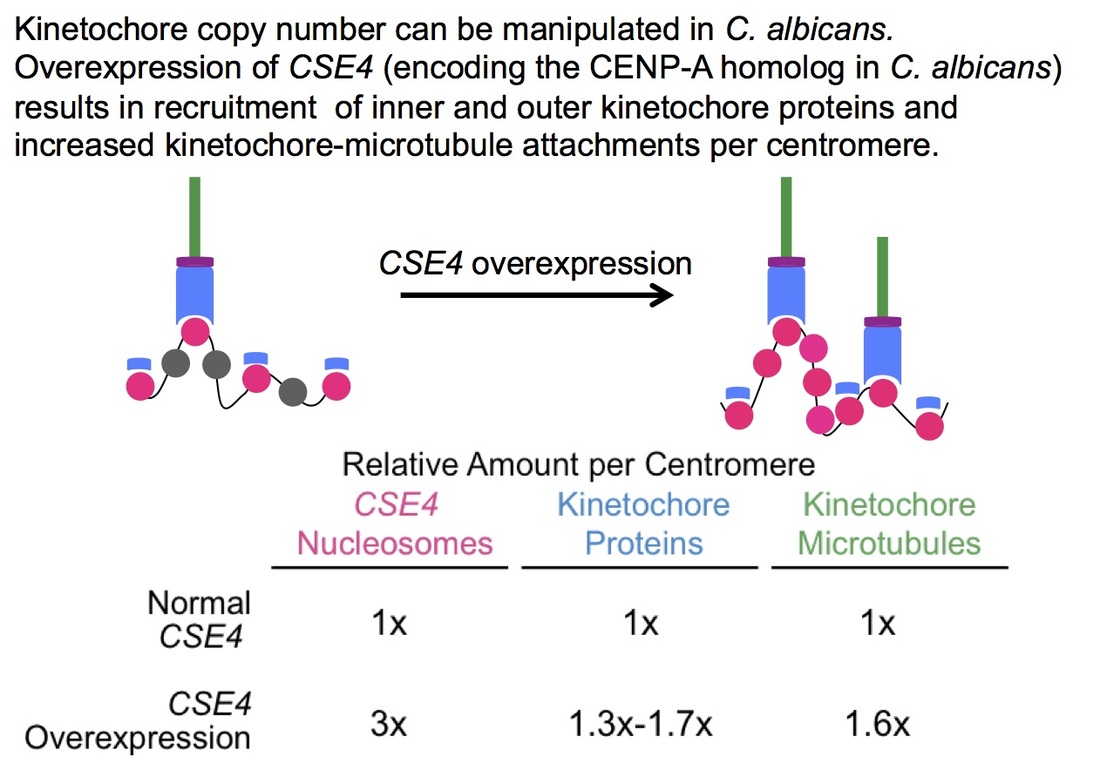
One of the main themes of my research is how cells can adapt the size and position of the kinetochore in response to stress or changes in the cellular physiology while maintaining viability and the ability to segregate chromosomes. Flexibility of kinetochore size and position is associated with human cancers. Overexpression of kinetochore proteins can alter the copy number of kinetochore complexes, can interfere with kinetochore function, and overexpression of kinetochore proteins in tumors correlates with poor patient prognosis. We have developed a model system in which kinetochore copy number can be manipulated in C. albicans to increase the number of kinetochore-microtubule attachments per centromere.
|
|
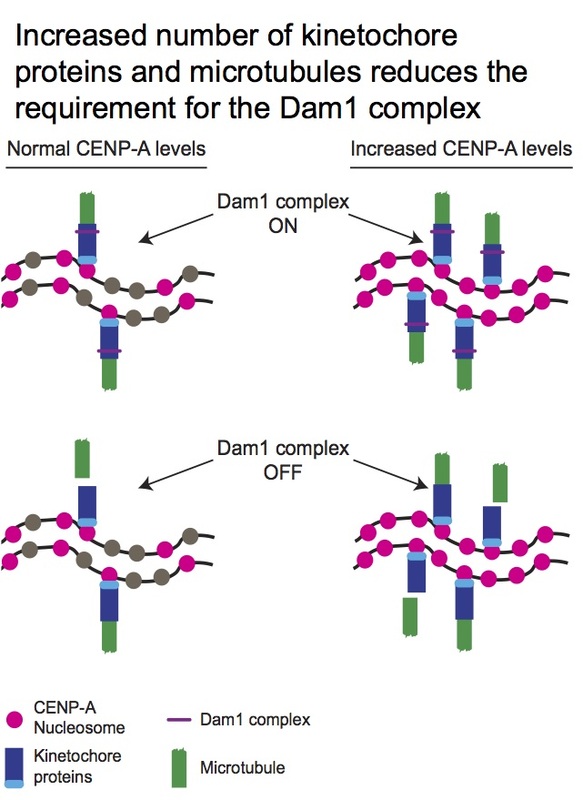
Using this model system, we addressed the question of why the Dam1 complex, a complex of fungal-specific kinetochore proteins important for processive attachment to microtubules, is required in some organisms, such as Saccharomyces cerevisiae and C. albicans, with one microtubule attachment per centromere, but not required in other fungi, such as Schizosaccharomyces pombe, with multiple microtubule attachments per centromere. We found that by increasing kinetochore copy number, Dam1 complex members became less critical for viability. Our results support the hypothesis that the number of microtubule attachments per centromere determines the requirement of the Dam1 complex and imply that the Dam1 complex functions as a processivity factor in vivo. This work was published in Current Biology and recommended in the F1000 database. We also used C. albicans to explore the role of the monopolin complex in promoting accurate chromosome segregation. We found that the primary role of the monopolin complex is to recruit condensin to organize centromere and repetitive DNA sequences. This manuscript has recently been published in Molecular Biology of the Cell and was selected for “Highlights from MBoC” in the ASCB newsletter.
|
|
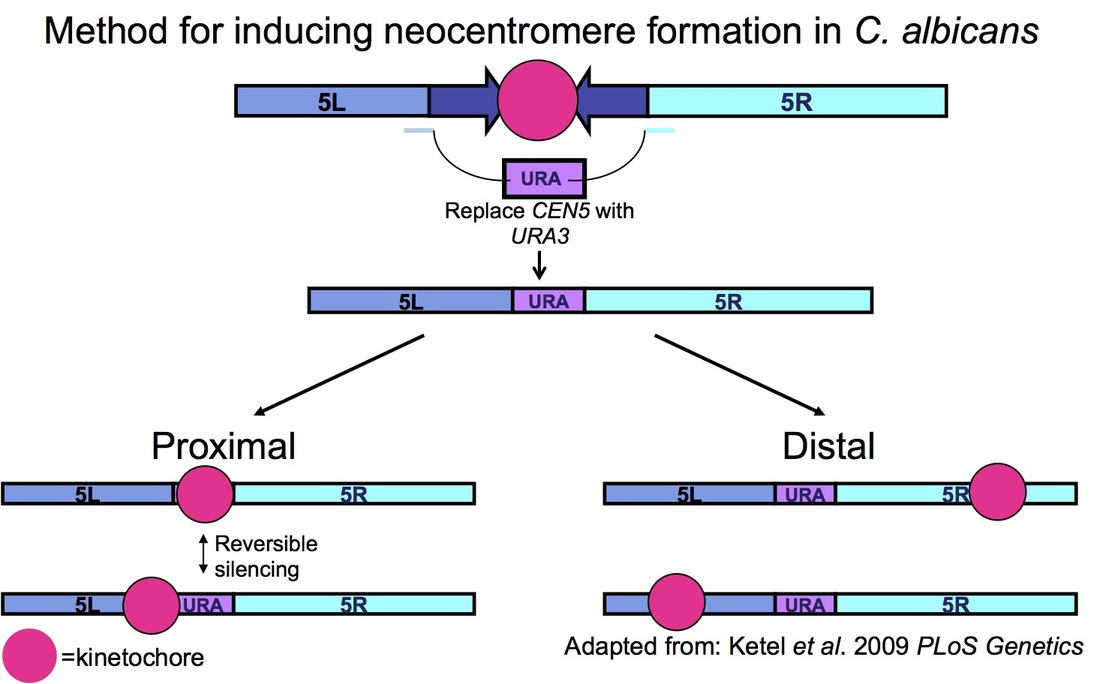
Neocentromeres, assembly of ectopic kinetochores following disruption of the native centromere, are associated with several types of cancer as well as with developmental disorders and birth defects. Deletion of C. albicans centromere DNA results in the efficient formation of neocentromeres either proximal to the native centromere or at distal loci on the chromosome arms, similar to what has been observed for human neocentromeres. To date, we have identified 20 neocentromere positions on C. albicans chromosome 5. Interestingly, these neocentromeres exhibit a range of chromosome stability rates, and some neocentromeres show high chromosome loss rates under stress, consistent with studies showing that human neocentromere has decreased error correction levels. Neocentromere positions also move following growth in stressful conditions indicating that neocentromeres show reduced inheritance compared to native centromeres. Current work focuses on eliciting the mechanisms of neocentromere function and inheritance and characterizing the evolutionary consequences of centromere alterations.
|
Publications related to current projects:
Anderson, M.Z., Wigen, L.J., Burrack, L.S., and Berman, J. (2015) Real-Time Evolution of a Subtelomeric Gene Family in Candida albicans. Genetics 200(3): 907-919. (click here for article)
Tsai, H-J., Baller, J.A., Liachko, I., Koren, A., Burrack, L.S., Hickman, M.A., Thevandavakkam, M.A., Rusche, L.N., and Berman, J. (2014) ORC binding, nucleosome depletion patterns and a primary sequence motif can predict origins of replication in a genome with epigenetic centromeres. mBio. 5(5):e01703-14. (click here for article)
Burrack, L.S., Clancey, S.E.A., Chacón, J., Gardner, M.K., and Berman, J. (2013) Monopolin recruits condensin to organize centromere DNA and repetitive DNA sequences. Mol Biol Cell. 24(18): 2807-2819. (click here for article)
Burrack, L.S. and Berman, J. (2012) Neocentromeres and epigenetically inherited features of centromeres. Chromosome Res. 20(5): 607-619. (click here for article)
Burrack, L.S. and Berman, J. (2012) Flexibility of centromere and kinetochore structures. Trends Genet. 28(5): 204-212. (click here for article)
Burrack, L.S., Applen, S.E., and Berman, J. (2011) The Requirement for the Dam1 Complex Is Dependent upon the Number of Kinetochore Proteins and Microtubules. Curr Biol. 21(10): 889-896. (click here for article)
Roy, B., Burrack, L.S., Lone, M.A., Berman, J. and Sanyal, K. (2011) CaMtw1, a member of the evolutionarily conserved Mis12 kinetochore protein family, is required for efficient inner kinetochore assembly in the pathogenic yeast Candida albicans. Mol Microbiol. 80: 14-32. (click here for article)
Koren, A., Tsai, H.J., Tirosh, I., Burrack, L.S., Barkai, N., and Berman, J. (2010) Epigenetically-inherited centromere and neocentromere DNA replicates earliest in S-phase. PLoS Genet. 6: e1001068. (click here for article)
Anderson, M.Z., Wigen, L.J., Burrack, L.S., and Berman, J. (2015) Real-Time Evolution of a Subtelomeric Gene Family in Candida albicans. Genetics 200(3): 907-919. (click here for article)
Tsai, H-J., Baller, J.A., Liachko, I., Koren, A., Burrack, L.S., Hickman, M.A., Thevandavakkam, M.A., Rusche, L.N., and Berman, J. (2014) ORC binding, nucleosome depletion patterns and a primary sequence motif can predict origins of replication in a genome with epigenetic centromeres. mBio. 5(5):e01703-14. (click here for article)
Burrack, L.S., Clancey, S.E.A., Chacón, J., Gardner, M.K., and Berman, J. (2013) Monopolin recruits condensin to organize centromere DNA and repetitive DNA sequences. Mol Biol Cell. 24(18): 2807-2819. (click here for article)
Burrack, L.S. and Berman, J. (2012) Neocentromeres and epigenetically inherited features of centromeres. Chromosome Res. 20(5): 607-619. (click here for article)
Burrack, L.S. and Berman, J. (2012) Flexibility of centromere and kinetochore structures. Trends Genet. 28(5): 204-212. (click here for article)
Burrack, L.S., Applen, S.E., and Berman, J. (2011) The Requirement for the Dam1 Complex Is Dependent upon the Number of Kinetochore Proteins and Microtubules. Curr Biol. 21(10): 889-896. (click here for article)
Roy, B., Burrack, L.S., Lone, M.A., Berman, J. and Sanyal, K. (2011) CaMtw1, a member of the evolutionarily conserved Mis12 kinetochore protein family, is required for efficient inner kinetochore assembly in the pathogenic yeast Candida albicans. Mol Microbiol. 80: 14-32. (click here for article)
Koren, A., Tsai, H.J., Tirosh, I., Burrack, L.S., Barkai, N., and Berman, J. (2010) Epigenetically-inherited centromere and neocentromere DNA replicates earliest in S-phase. PLoS Genet. 6: e1001068. (click here for article)
Funding for centromere and kinetochore research projects:
Grinnell College
2013 – 2015
Committee for Faculty Scholarship Funds Grant
Mentored Advanced Project for Undergraduate Training
American Cancer Society
2012 – 2013
Postdoctoral Fellowship
N.I.H. Ruth L. Kirschstein Individual National Research Service Award
2008 – 2011
Postdoctoral Fellowship
Grinnell College
2013 – 2015
Committee for Faculty Scholarship Funds Grant
Mentored Advanced Project for Undergraduate Training
American Cancer Society
2012 – 2013
Postdoctoral Fellowship
N.I.H. Ruth L. Kirschstein Individual National Research Service Award
2008 – 2011
Postdoctoral Fellowship
Recent Collaborators and Undergraduate Research Students:
Ph.D. Research:
|
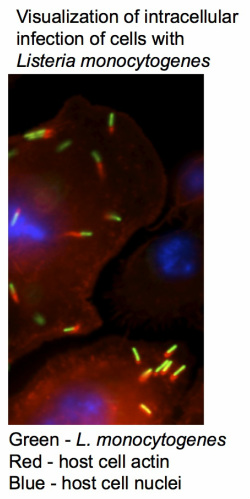
In my graduate work in Darren Higgins’ lab at Harvard University, I focused on host-pathogen interactions between the intracellular bacterial pathogen Listeria monocytogenes and its host cells. Upon entry into host cells, L. monocytogenes escapes the entry vacuole, replicates within the cytosol, and uses actin-based motility to spread to neighboring cells. The goal of my thesis research project was to identify and characterize host cell factors capable of modulating vacuole escape of L. monocytogenes. My thesis work began by using Drosophila as a model eukaryote to conduct a genome-wide RNA interference (RNAi) screen for host factors required for L. monocytogenes intracellular infection. The genome-wide RNAi screen of Drosophila cells identified host factors with diverse functions required for L. monocytogenes infection. Most striking was the large number of vesicular trafficking pathway components identified. I then extended the analysis to human tissue culture cells and confirmed many of the Drosophila screen results by developing a system to perform small-scale RNAi screens using shRNAs to target specific subsets of the genome. I also used RNAi in human cells to address an open question in the L. monocytogenes field. In the mouse model of infection, L. monocytogenes mutants that are unable to escape the phagocytic vacuole are avirulent. In murine cell lines and in the mouse model of infection, the pore-forming cytolysin listeriolysin O (LLO) is absolutely required for vacuole escape. However, in several different types of human cell lines, vacuole escape of LLO-negative (deltaLLO) L. monocytogenes has been observed, and the mechanism permitting this LLO-independent escape is not well understood. Using RNAi, I identified knockdowns that permitted increased LLO-independent vacuole escape in human cells. Follow-up studies examining vacuolar maturation following knockdown of the early endosome/recycling endosome regulator RABEP1, and work comparing vacuolar maturation in several human cell lines provided support for the hypothesis that LLO-independent vacuolar escape may be inversely correlated with the rates of vacuolar maturation.
|
Publications related to dissertation research:
Burrack, L.S., Harper, J.W., and Higgins, D.E. (2009) Perturbation of vacuolar maturation promotes listeriolysin O-independent vacuolar escape during Listeria monocytogenes infection of human cells. Cell Microbiol. 11: 1382-1398. (click here for article)
Burrack, L.S. and Higgins, D.E. (2007) Genomic approaches to understanding bacterial virulence. Curr Opin Microbiol. 10(1): 4-9.
Agaisse, H.*, Burrack, L.S.*, Philips, J.A., Rubin, E.J., Perrimon, N., and Higgins, D.E. (2005) Genome-wide RNAi screen for host factors required for intracellular bacterial infection. Science. 309: 1248-1251. (*These authors contributed equally.) (click here for article)
Gründling, A., Burrack, L.S., Bouwer, H.G., and Higgins, D.E. (2004) Listeria monocytogenes regulates flagellar motility gene expression through MogR, a transcriptional repressor required for virulence. Proc Natl Acad Sci USA. 101: 12318-12323. (click here for article)
Burrack, L.S., Harper, J.W., and Higgins, D.E. (2009) Perturbation of vacuolar maturation promotes listeriolysin O-independent vacuolar escape during Listeria monocytogenes infection of human cells. Cell Microbiol. 11: 1382-1398. (click here for article)
Burrack, L.S. and Higgins, D.E. (2007) Genomic approaches to understanding bacterial virulence. Curr Opin Microbiol. 10(1): 4-9.
Agaisse, H.*, Burrack, L.S.*, Philips, J.A., Rubin, E.J., Perrimon, N., and Higgins, D.E. (2005) Genome-wide RNAi screen for host factors required for intracellular bacterial infection. Science. 309: 1248-1251. (*These authors contributed equally.) (click here for article)
Gründling, A., Burrack, L.S., Bouwer, H.G., and Higgins, D.E. (2004) Listeria monocytogenes regulates flagellar motility gene expression through MogR, a transcriptional repressor required for virulence. Proc Natl Acad Sci USA. 101: 12318-12323. (click here for article)
Funding for dissertation research:
Howard Hughes Medical Institute
2002 – 2007
Pre-doctoral Fellowship
Howard Hughes Medical Institute
2002 – 2007
Pre-doctoral Fellowship